Contents
Cell Biology and Genetics
Physiology and Pharmacology
Cell Organelles
Cells are the basic unit of our bodies. They’re all made up of subcellular components called organelles. The type of organelles present differ cell to cell but there are some general organelles that you will find in most cells. We will look at these organelles.
Key organelles:
- Nucleus
- Rough endoplasmic reticulum
- Smooth endoplasmic reticulum
- Ribosomes
- Mitochondria
- Lysosomes
- Golgi apparatus
- Peroxisomes
- Cell-surface membrane
- Cytoskeleton
Nucleus:
Function:
The nucleus is the largest organelle in a cell. It’s often described as being the control centre of the cell. It coordinates the cell’s activities such as growth, metabolism, protein synthesis and cell division. It’s where most of the DNA, nuclear proteins and RNA is stored.
Structure:
- Nucleolus- site of rRNA synthesis
- Appears as a black circle within the centre of the nucleus
- Nuclear envelope- Double membrane envelope
- Acts as a partially permeable membrane
- Nuclear pore- Microscopic pores in the nuclear envelope
- Allows macromolecules to pass through
- Nucleoplasm-Type of protoplasm which functions in serving as a suspending substance in the nucleus + aids in the maintenance of structure and shape of the nucleus
- DNA in nucleus appears in 2 forms: Heterochromatin and Euchromatin
- Heterochromatin – Appears darker- DNA more condensed results in less RNA synthesis
- Euchromatin – Appears lighter- DNA less condensed results in more RNA synthesis
Rough Endoplasmic reticulum (RER):
Function:
The rough endoplasmic reticulum is usually continuous with the nuclear envelope. It functions in intracellular transport and the synthesis of proteins. The proteins produced are destined for extracellular transport, lysozymes or cell membranes
Structure:
- Consists of series of connected flattened sacs (cisternae)
- Ribosomes are continuous with the outer layer of the cisternae
- Also has vesicles that bud off from it
Smooth Endoplasmic Reticulum (SER):
Function:
The smooth endoplasmic reticulum is responsible for lipid biosynthesis, steroid formation and intracellular transport
Structure:
- Consists of series of connected flattened sacs (cisternae-less flat than RER and more tubular)
- Also has vesicles that bud off from it
Ribosomes:
Function:
Functions in protein synthesis
Structure:
Ribosomes are composed of 2 rRNA subunits (which are integrated with proteins) that wrap around mRNA to initiate the translation section for protein synthesis.
Lysosomes:
Function:
Lysosomes break worn out organelles and digest pathogens. They can also ingest and digest bacteria and viruses that enter the cell.
Structure:
Membrane bound organelle that contains lysozymes + acid hydrolases (protease, lipase, glycolipase etc)
Golgi Apparatus:
Function:
Packaging and modifying proteins
Structure:
- Crescent-like shaped cisternae
- Vesicles from RER and fuse with convex side of Golgi
- Proteins move from convex to concave (trans to cis)
- Vesicles bud off from concave side
Peroxisomes:
Function:
- Chemical detoxification and lipid biosynthesis
- Ingest toxins
- Contains enzymes that remove and transfer hydrogen atoms from toxins to water to create hydrogen peroxide.
- Enzymes in peroxisomes turn the hydrogen peroxide into water and oxygen.
Structure:
Peroxisomes are small vesicles, single membrane bound organelle that contains digestive enzymes.
Mitochondria:
Function:
The function of the mitochondria is for ATP synthesis for energy production
Structure:
Mitochondria have a double membrane with an outer and inner lipid bilayer. The inner layer is significantly larger than the outer one hence it becomes highly folded into structures called cristae. Within the inner membrane lies the matrix which holds the mitochondrial: DNA, ribosomes and enzymes. The function of the mitochondria is for ATP synthesis for energy production
Cell-surface membrane:
Functions:
- Semi-permeable- Allows entry of small non-polar molecules
- Compartmentalisation of cells and organelles
- Communication
- Recognition
- Electrical/chemical stimulation in response to stimuli
Structure:
- Surrounds the cell, acts as a barrier
- Composed of phospholipid bilayer with embedded proteins attached (Fluid Mosaic model)
- Cholesterol embedded in bilayer – controls fluidity + rigidity
Cytoskeleton:
Functions:
- Responsible for maintaining and changing cell shape. It provides structural support for the plasma membrane + cell organelles and it also allows the movement of organelles and substances released from them around the cytosol and to different areas of the cell
- This organelle allows contractility in cells such as those in the muscle, and also plays a key role in the cytokinesis stage of mitosis.
Structure:
The cytoskeleton consists of microtubules, microfilaments and intermediate filaments.
DNA Structure and Replication:
- The genetic code of life (all cellular organisms and most viruses)
- Contains the genetic material of the cell
- Found in nucleus in the cell
- Genetic code is almost universal
Structure of DNA:
Monomer of DNA has 3 parts:
| Components | Composition |
| 1)Pentose sugar (2-Deoxyribose) 2)Phosphate group 3)Nitrogenous base | Pentose sugar + phosphate group form the sugar phosphate backbone via phosphodiester bonds |
A-adenine,T-thymine,G-guanine,C-cytosine – Bases in DNA
- Purine=A,G (Double-ring)
- Pyrimidine=C,T (Single-ring)
DNA is a double stranded molecule. The 2 strands are complementary to each other and anti-parallel. Double strand twisted to form right-handed double helix.
Double helix contains major and minor grooves. It is easier for proteins to form complexes on major grooves as they are more exposed
Complimentary base pairing:
- A goes to T (2 hydrogen bonds)
- C goes to G (3 hydrogen bonds)
Central Dogma Of Biology
DNA → RNA → Protein
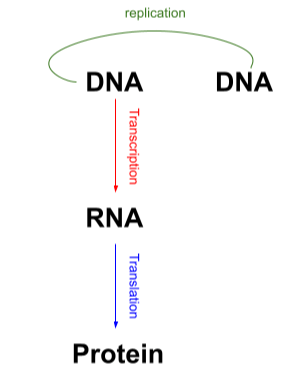
The 3 models of DNA Replication:
Conservative replication- The original DNA molecule is used as a template and the double helix stays intact
Results in: One original parent molecule + one completely new daughter double helix
Semi-conservative replication- The original DNA double helix is broken up into 2 single strands which are then used as templates for new complementary bases to form the complementary strand
Results in: 2 DNA strands produced each strand conserving half the DNA from the first strand
Dispersive model-Parental double helix is broken into double helix segments which acts as templates for new DNA synthesis. Segments then join together.
Results in: 2 DNA strands that are mixtures of parental and daughter DNA.
DNA Polymerase
DNAPs in E.Coli
- DNAP I: Functions in repair and replication
- DNAP II: Functions in repair
- DNAP II: Principle DNA replication enzyme
- DNAP IV: Functions in DNA repair
- DNAP V: Functions in DNA repair
3 actions of DNA polymerase:-
- 3’ to 5’ exonuclease activity
- 5’ to 3’ exonuclease activity
- 5’ to 3’ polymerising activity
Exonucleases:
Exonucleases are enzymes that removes successive nucleotide bases from the end of a polynucleotide strand
3’ to 5’ exonuclease activity-
- Addition of an incorrect base stalls the action of DNA polymerases-it can’t add the next nucleotide in the sequence
- Exonucleases removes 1 nucleotide base at a time
- Polymerase reversing direction (moving from synthesising 5’ to 3’ to 3’ to 5’ and correcting a single base mistake)
- Proof-reading is quite a slow process
5’ to 3’ exonuclease activity-
- Removes upto 10 nucleotide bases at a time
- Removes the RNA primer at the end of replication
DNA POL1:
- Exonuclease activity used as a means of proofreading
- Removes incorrect bases and replaces them with correct ones
- Also degrades the RNA primer and replaces it with a newly synthesised strand of DNA at the end of replication
- Also has some replication capabilities but is too slow (600 dNTPs added/minute) and moderately processive (refers to the amount of dNTPs that can be added until the enzyme falls off) to be the principal replication enzyme
- Functions in multiple processes that require short lengths of DNA segments
DNA POL2:
- Also participates in exonuclease activity
- Similar to POL1
DNA POL3 has synthesis activity:
- Main synthesis molecule in DNA replication
- Synthesises 5’ to 3’ direction
- Fast (1000 dNTPs added/sec), highly accurate plus highly processive (can add a lot of nucleotides at a time without dissociating)
DNA Replication:
Split into 3 parts:
- Initiation
- Elongation
- Termination
Initiation:
- DNA Helicase breaks the hydrogen bonds between the 2 strands to unwind the double helix at replication origins which are rich with A-T pairs
- RNA Primase catalyses the synthesis of a short RNA segment upfield of the DNA polymerase. These short segments are called primers.
- Primers help to expose the 3’ end of DNA so DNA polymerase can bind to the strand and begin synthesising
Elongation:
- DNA polymerase III binds to the 3’ end of the primer. Makes a continuous strand of DNA on the 3’ to 5’ strand (synthesising 5’ to 3’)
- 5’ to 3’ strand is referred to as “lagging strand” as helicase unwinds the DNA strand, polymerase has to join on the 3’ end leading to multiple polymerases synthesizing the new strand resulting in okazaki fragments
- Before polymerase can bind to the DNA strand RNA primase must synthesise more primers allowing the binding of polymerases
- Okazaki fragments have to be adjoined by DNA ligase
- DNA Topoisomerase helps keep the original DNA strands unwound so the DNA polymerase can continue to synthesise
Termination:
- The primers (segments of RNA) present in the newly synthesised strand are stripped away by exonucleases which polymerases replace with DNA
- More than one helicase acts on DNA when replication takes place, so there are multiple replication forks
- The 2 okazaki fragments at the place of two forks meeting are all joined by DNA ligase to form 2 continuous DNA molecules (Semi-conservative)
DNA Packaging:
DNA is wrapped twice around histone octomer proteins forming nucleosomes in a ‘beads on a string’ formation. Nucleosomes further foiled into solenoids. Solenoids can further loop and coil into chromosomes.
Total of 10 Histone isoforms:
- 7 somatic
- 3 germline
Chromatin:
- Chromatin-complex of DNA and histones
- 2 types of chromatin-Heterochromatin and Euchromatin
- Heterochromatin– Highly condensed therefore appears darker
Lacks RNA synthesis
- Euchromatin– Less condensed therefore appears lighter
Has RNA synthesis
Telomeres:
- Telomeres are the end regions of chromosomes
- Clusters of TTAGGG- can be thousands of base pairs long
- Help protect the integrity of chromosomes + allows replication of extreme ends
- Telomeres slowly eaten away after every division
- Makes the replicated strands shorter as the telomere sequence signals DNA polymerase to unbind and leave the strand. Therefore as replication occurs strands get shorter
- Related to aging
Cell cycle:
- Cell cycle is the sequence of events leading up to and during the process of cell division.
- Somatic cells- all the cells in the body that aren’t haploid / gamete cells. Diploid with the full set of 46 chromosomes.
- Two main components of the cell cycle:
- Interphase: Cell prepares to divide.
- Mitosis: Actual process of cell division.
- Interphase:
3 main subphases.
- G1: This is when the cell continues to grow and replicate organelles, synthesising proteins ready for DNA replication.
- S: DNA replication. 2 complete copies of DNA are available.
- G2: cell increases in size and produces more proteins. Checks if cell is ready for mitosis
- M phase:
2 main components:
- Cell Division
- Cytokinesis
- After the mitosis phase cells enter G1 and continue to divide once again.
- Some cells can stay in a “rest phase” : G0. This is where the cell does not divide or prepare to divide. This is common in skeletal cells and neurons.
Chromosomes
- Not visible at interphase. They condense at the mitotic phase – during prophase.
- Post replication they contain 2x the normal amount of chromatids.
- P arm → short arm
- Q arm → long arm
- Healthy somatic contain 46 chromosomes – 23 pairs. 1 maternal set and the other is a paternal set.
- Gametes are haploid – they contain a single set of chromosomes (23 chromosomes).
- Homologous chromosomes: have the same gene loci (same region on the chromosome) but different alleles
Karyotyping
- Method of showing all chromosomes under a light microscope.
- Looks for number of chromosomes, position of centromeres on chromosomes, sex determining, lebgth of chromosomes,
- Chromosomes are taken during the metaphase stage of mitosis. Condensed chromosomes all line up on the metaphase plate. All chromosomes are visible.
Mitosis:
- Single cell divides into two daughter cells, identical to the parent cell and includes the complete genome identical.
- Important for:
- development of embryos
- Growth
- Healing
- Repair of damage
- Replacement of damaged cells.
- Stages:
Prophase:
- Centromeres holding the chromatids together are surrounded by a kinetochore.
- Chromatin fibres condense and shorten to form chromosomes.
- Centrioles move towards the poles (Responsible for spindle fibre production of cells)
- Centrosomes consist of 2 centrioles perpendicular towards each other.
Prometaphase:
- Spindle fibres of centrioles will extend and connect to the kinetochore of the chromosomes.
- Nuclear membrane disintegrates, causing the chromosomes to lie free within the cell.
Metaphase:
- Chromatids are led by mitotic spindle fibres
- Chromatids line up on the metaphase equatorial plate
Anaphase:
- Centromere and kinetochore split. Allows chromosomes to separate into individual sister chromatids.
- Microfibres shorten and centrosomes will pull with equal force at opposite poles, allowing full separation of chromatids.
Telophase:
- Nuclear envelope reforms
- Nucleolus reappears
- Chromosomes uncoil into chromatin (no longer so condensed)
- Microtubules de-attach from kinetochore
Cytokinesis:
- Begins halfway through anaphase
- Results in the splitting of parent cell into 2 identical daughter cells.
- Late anaphase: contractile ring develops along line previously occupied by the metaphase plate.
- Allows the formation of the “cleavage furrow”, stimulates the pinching of the cytoplasm
- As cleavage furrow deepens the cells will eventually separate with own nucleus, own chromosomes and cytoplasm
Meiosis:
- Occurs in germ line cells – produces gametes.
- End product is 4 non-identical cells with half the chromosome content of a somatic cell (haploid, 23 chromosomes)
- Cells don’t go through G2 phase – stop at S phase of the interphase
- 2 steps
- Meiosis I
- Meiosis II
- Stages
Prophase I:
- Chromatin Fibres condense and shorten
- Microtubules extend from poles to the centre of the cell.
- Spindle microtubules attach to the kinetochores of each chromosome from the centrosomes
- Nuclear envelope breaks up and nucleolus diminishes
- Crossing Over
- Exchange of genetic material between homologous chromosomes. Results in recombinant chromosomes during sexual reproduction
- Occurs between prophase I and metaphase I
- Allows genetic variation to occur. Chromatids are held together by the centromere. No longer identical hence after separation daughter cells will continue a combination of both maternal and paternal alleles.
Metaphase I:
– Homologous pairs will line up along the metaphase plate.
Anaphase I:
– Microtubules attached to each kinetochore shorten.
-Centrioles pull with equal force from opposite poles
-Allow 1 complete chromosome from each homologous pair to be pulled towards each pole.
Telophase I:
– Nuclear Envelope reforms
– Microtubules of the mitotic spindle break up
-Chromatin uncoils and becomes looser as the nuclear membrane is formed
Cytokinesis I:
-Cleavage furrow and causes the separation of the cytoplasm
6. Meiosis II:
-Exact same steps as mitosis.
- Interphase of Meiosis I: 46 x 2 chromosomes = 92 Chromatids
- End of Meiosis I: 46 Chromatids (genetically variated)
2 cells
- End of Meiosis: 23 Chromosomes
4 cells
- 1 diploid germ cell divided into 4 gametes in males, all ae functional. Female ovum: ¼ of the gametes are functional cells. Other 3 are resulting daughter cells degenerate polar bodies
Gamete Formation
Male Gamete Formation
- Sperm are produced after puberty
- Production of sperm is continuous and prolific. Each ejaculation contains 100-650 million sperm cells.
- Occurs within the testes
- Spermatogonia are the stem cells that give rise to sperm. Located in the periphery of the seminiferous tubule
- Developing sperm move towards the central opening as they undergo meiosis and differentiation
- 1 spermatogonia = 4 spermatozoa. Spermatogonia is the diploid stem cell. Sperm cells are haploid.
Female gamete formation
- Development of ova (unfertilised, mature egg) occurs within the ovary.
- Oogonia are the stem cells that give rise to ova. Multiply and begin meiosis.
- Stops at prophase 1 – here ova are primary oocytes.
- Puberty causes stimulation of FSH and LH
- FSH and LH regulate development, growth, pubertal maturation, reproductive processes of the female body.
- FSH and LH is synthesised and secreted by gonadotropic cells of the anterior pituitary gland
- FSH periodically stimulates the growth of a follicle and induces the primary oocyte to complete meiosis I and start meiosis II. This is a secondary oocyte
- Halt of meiosis II in oocytes. Continues after the oocyte has been penetrated by a sperm cell , where LH stimulates completion of meiosis II.
- Birth of a female: no more oocytogonium, only primary oocytes.
- 14th day of the menstrual cycle: primary oocyte turns into a secondary oocyte. Stops at metaphase II till sperm penetration. If fertilisation occurs a zygote is formed, if not degeneration occurs.
- 1 reproductive cell and 3 polar bodies. Egg needs nutrients to support life so excess chromosomes are discarded as one section takes everything it needs.
- Nutrients are distributed to only one cell. This helps to support ovulation
Mechanisms for mutations in the sperm and egg:
- Women don’t have many germ-line cells. They start with a pool of egg cells that degenerate from puberty. 1 degenerates each month if no fertilisation occurs.
- Older eggs are more susceptible to mutations and destruction. This is because they are exposed to more risk factors caused by life-style and environment that can potentially mutate the egg.
- Spindle assembly mitotic checkpoint occurs just after the metaphase before the anaphase. This checkpoint ensures all chromosomes are lined up by spindles on the metaphase plate.
- Inhibits progression of mitosis to anaphase until all homologous chromosomes are aligned to the spindle apparatus.
- Unequal separation of chromosomes can lead to trisomy and other disorders
- Lifestyle and environmental factors can increase the risk of aneuploidy – presence of an abnormal number of cells.
- Most aneuploidy are maternally derived. This is because of prolonged arrest of oocytes causing reduced activity of the SAC, leading to maternal age linked errors in segregation control.
Genetic Disorders
Laws of segregation:
- Allele pairs separate during gamete formation but unite during fertilization
- Aneuploidy = an abnormal amount of chromosomes due to improper segregatio post anaphase
- Trisomies = additional chromosome leading to 3 homologous chromosomes at a particular chromosome instead of 2.
(2n+1)
- Monosomies is 1 missing chromosomes. This means there is 1 chromosome instead of a homologous pair.
(2n-1)
- Polypody = the total number of chromosomes is a multiple other than 2 (e.g. 3n, 4n, 5n where n=23 chromosomes)
- Nondisjunction is the failure of chromosome separation in anaphase of mitosis or meiosis.
- Results in a 1:1 ratio of daughter cells with an extra chromosome (2n+1) to those with a loss of a chromosome (2n-1)
- If occur during meiosis it is a germ-line mutation and therefore transmissible to the next generation
- In Meiosis I : failure of homologous chromosomes to separate
- In Meiosis II: Failure of sister chromatids to split
- Nondisjunction occurs due to spindle fibres not attaching properly leading to unequal splitting of chromosomes
Disorders:
- Polyploidy – an uneven gain of a whole haploid set of chromosomes.
- Caused by 2 sperms fertilising the same egg. Zona pellucida fails to form a hard protective layer
- Incorrect meiotic genesis leading to diploid copies of chromosomes within single oocyte. Diploid oocyte + haploid sperm = triploidy
- Trisomy 21: Down’s Syndrome
– Autosomal, 3 copies of chromosome 21
– Results in varying degrees of mental disability, decreased immunity to diseases and organ defects. Distinguishable facial features. Increased prevalence of leukaemia. Increased incidence of early Alzheimer’s disease
- Trisomy 18: Edward’s Syndrome
– Autosomal: 3 copies of chromosome 18
-Failure of all organ systems, death in a few months, small lower aw, low set ears, overlapping fingers
- Trisomy 13: Patau’s Syndrome
– Caused by 3 copies of chromosome 13
-Multiple congenital abnormalities: polydactyly (more than 5 fingers/toes), holoprosencephaly (prosencephalon fails to develop within the two hemispheres), die within a month.
- Monosomy 45: Turner’s Syndrome
– Only 1 X chromosome being inherited
-Results in a sterile female, short in stature, heart and kidney defects, reduced effects of puberty
- Trisomy 47: Superfemale
– 3 X chromosomes inherited.
– Results in healthy fertile female, delayed motor development, low IQ, delayed speech, abdominal pain
- Trisomy 47: Klienfelter’s Syndrome
– Extra X chromosome in male (XXY)
-Sterile male: taller and less muscular, broader hips, longer legs, larger breasts, weaker bones, small penis and testicles, delay in puberty and less facial and body hair post puberty.
- Trisomy 47: Klienfelter’s Syndrome
-Sex chromosome inherits 2 Ys (XYY)
-Results in a sterile male, violent, decreased intelligence and taller male.
Non-disjunction of sister chromatids
- If nondisjunction occurs during mitosis, after ovum and sperm fuse (post-zygotic), individual will exhibit mosaicism.
- Mosaicism: the state of being composed of cells of two genetically different types.
- Earlier the mutation that occurs during embryogenesis → greater the number of aneuploid fetal cells
- Degree of mosaicism depends on when the mitotic error has occurred.
Translation
- Translation is the last process in the central dogma of biology- the formation of proteins. It’s the synthesis of proteins using mRNA and ribosomes.
- Genetic information from gene to protein is based on triplet code (codon). A codon is a triplet of 3 mRNA bases. Each codon codes for a particular amino acid.
- Codons read 5’ to 3’ direction
- Components of translation: mRNA, tRNA, 2 ribosomal subunits
The tRNA molecule:
- tRNA used bring amino acids that are complementary to the mRNA codon.
- tRNA transfers amino-acids to a growing polypeptide chain
- Each tRNA contains an anticodon which is complimentary to the mRNA strand. This binding ensures that the right amino acid has been brought for the codon.
- The tRNA molecule consists of a single RNA strand approximately 80 nucleotides in length. This molecule is clover shaped.
- L shaped -5’ end and 3’ end exist on the same side of the structure
- 3’ end of the tRNA acts like the amino-acid attachment site
Accurate translation:
- Aminoacyl-tRNA synthetase-ensures correct match between an amino-acid and tRNA. The reaction is from ATP -> AMP + 2Pi
- Correct match between tRNA anticodon and mRNA
Flexible pairing at the third base of a codon-wobble
- Allows some tRNAs to bind to more than one codon, allowing more flexibility at the third base of a codon – meaning binding more relaxed.
Ribosome structure:
- Facilitate specific coupling between tRNA anticodons with mRNA codons
- Subunits of ribosome made up of rRNA and protein
- 2 Subunits of ribosomes (small subunit and large subunit)
- 3 sites on a ribosome: E site, P site and A site (Think APE)
- P site holds tRNA that carries the growing polypeptide chain
- A site holds the tRNA that carries the single amino-acid to be added to the chain
- E site is the exit site, where tRNAs leave the ribosomes
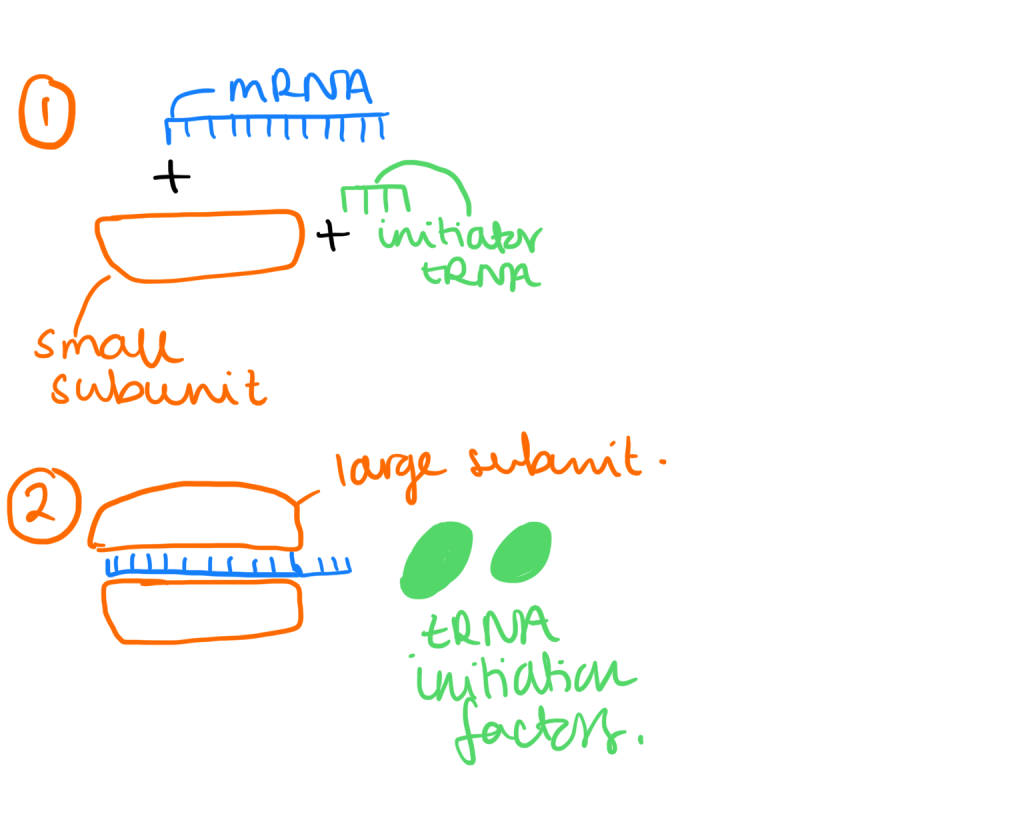
Initiation of translation:
- Small ribosomal subunit binds with mRNA and initiator tRNA (which carries methionine)
- Small subunit moves along the mRNA until it comes up to the start codon (AUG). Then proteins called initiation factors bring the large subunit and completes the translation initiation complex
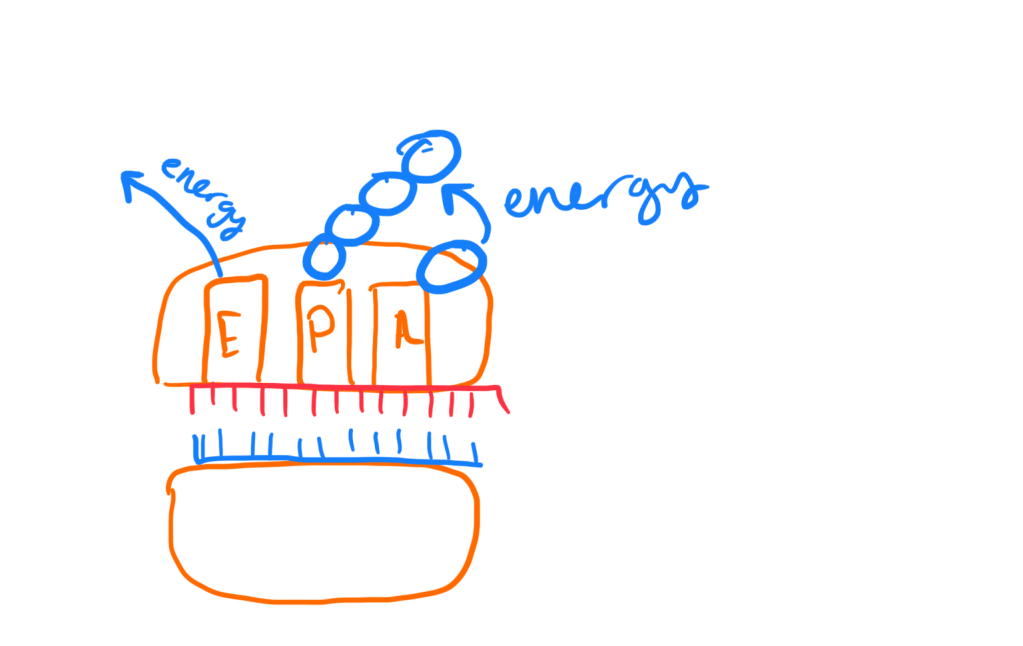
Elongation of polypeptide chain:
- Amino-acids added one at a time to the C-terminus of the growing polypeptide chain (C-terminus has exposed carboxyl end). This is the 3′ end
- Each addition of an amino acid utilises elongation factor proteins
- Step 1: Codon recognition
- tRNA with associated anticodon is present on the A site of the ribosome
- Step 2: Peptide bond formation
- Formation of a peptide bond between an amino and carboxyl groups of adjacent amino acids in polypeptide chain. The growing polypeptide chain from the tRNA in the P site is transferred to the single amino acid on the A site. This process requires an energy source via GTP
- mRNA shifts exactly by one codon on the ribosome. This process requires energy. The tRNA in the P position then shifts to the E site where it leaves the ribosomes and the tRNA in the A site occupies the P site.
- When a new tRNA binds to the A site this requires energy: GTP -> GDP + Pi
- When a tRNA leaves the E site of the ribosome, this also requires energy: GTP -> GDP+Pi
3. Termination of translation:
- Elongation of the polypeptide chain continues until the presence of the stop codon on the A site of the ribosome
- The A site accepts a release factor which causes the addition of a water molecule instead of an amino-acid.
- The addition of water releases the polypeptide chain and the translation complex all dissociate from each other. (Energy input is needed for this to occur)
- Release factor hydrolyses the mRNA chain and allows the release of the polypeptide chain from the tRNA in the P site
- This causes the whole translation complex to break away. Energy is required for this process: 2GTP -> 2GDP + 2Pi
Polyribosomes:
- Multiple ribosomes translating a single mRNA
- Allows faster and more efficient synthesis of a protein
- Each ribosome translates the same mRNA and makes the same proteins (useful because the cell might need multiple copies of the same protein). This is because cells need multiple copies of the same protein.
- Translation in bacteria occurs immediately after translation without post transcriptional modifications.
Polypepetide Targetting:
- Ribosomes can be of two types: free ribosomes and bound ribosomes.
- Free ribosomes: mostly synthesise proteins that function in the cytosol.
- Bound ribosomes: synthesise proteins that are part of the endomembrane system and that are secreted outside of the cell.
- Ribosomes can switch from free to bound.
- Protein synthesis may occur totally in the cytosol unless a signal is received by which it will be directed to the RER. This is marked by a signal polypeptide.
- The signal recognition particle will bind the the signal polypeptide, escorting the ribosome to attach to the RER.
- Proteins move to the correct locations of the cell through the specific peptide sequences. This could be inside the cytosol, the nucleus or the cell membrane.
Protein Structure and Protein Folding
Protein Function:
- Catalysts – enzymes
- Transporter – haemoglobin transports oxygen
- Structural support – collagen
- Immune system – antibodies and immunoglobin
- Ligands in cell signalling – hormones and neuotransmitters
- Ions channels and receptors – on the cell membrane
Protein Structure
- Proteins are polypeptides made by amino acids that are covalently bonded together, which forms a sequence.
- The amino acid is encoded by a gene. This means the nucleotide sequence of an protein determines sequence of amino acids
- The final polypeptide sequence folds into a highly specific 3D shaped protein, determined by the polypeptide sequence.
- Oligopeptides are short sequences of amino acids with a small sequence and a low amount of amino acid residues.
- Polypeptides are longer sequences of amino acids with a greater amount of amino acid residues
Chemical Structure of Amino Acids
- Amino acids are the monomers of polypeptide chains, which form proteins.
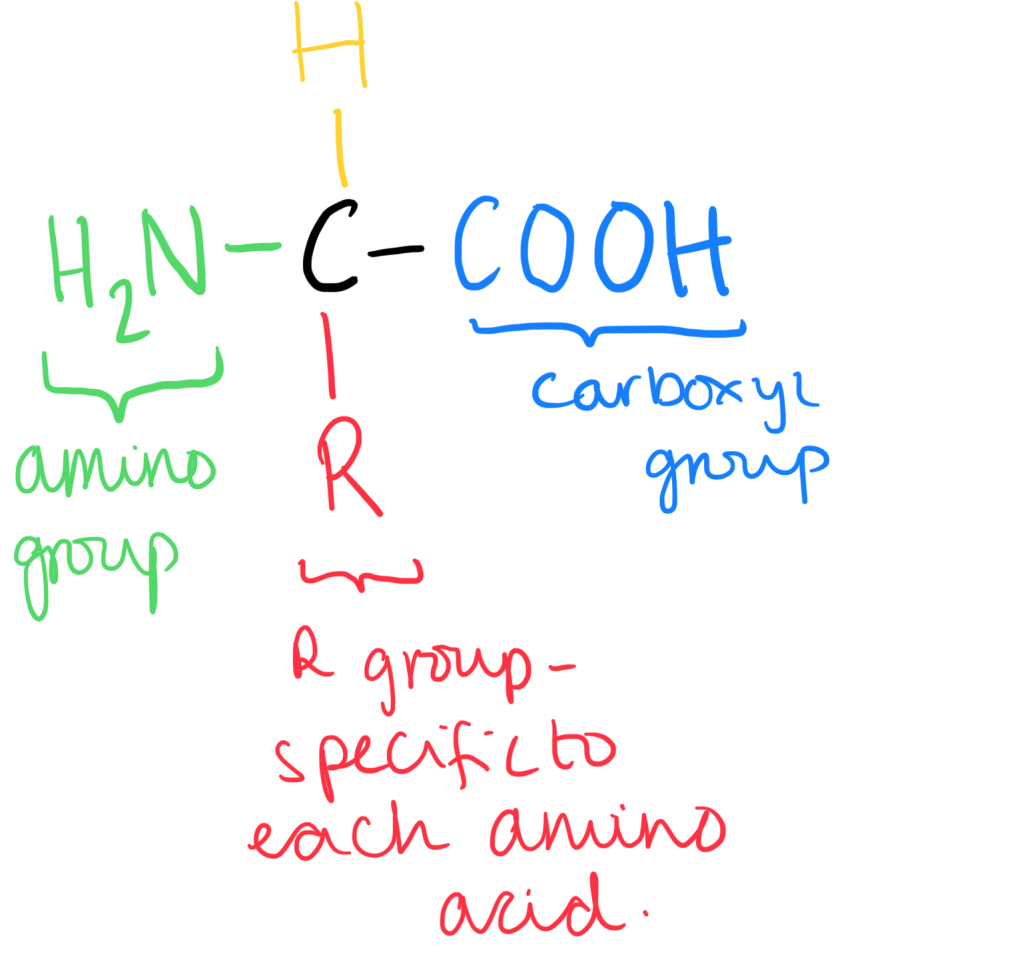
- Amino acids can become ionised as the amino group acts as a base (proton acceptor). They can accept the hydrogen ion from the carboxyl group.
- As the charged amino acid has a positive and a negative charge, the net charge of the amino acid is zero. This form is called a zwitterion.
- Depending on the pH of the solution, the ion can be positively charged cation or a negatively charged anion.

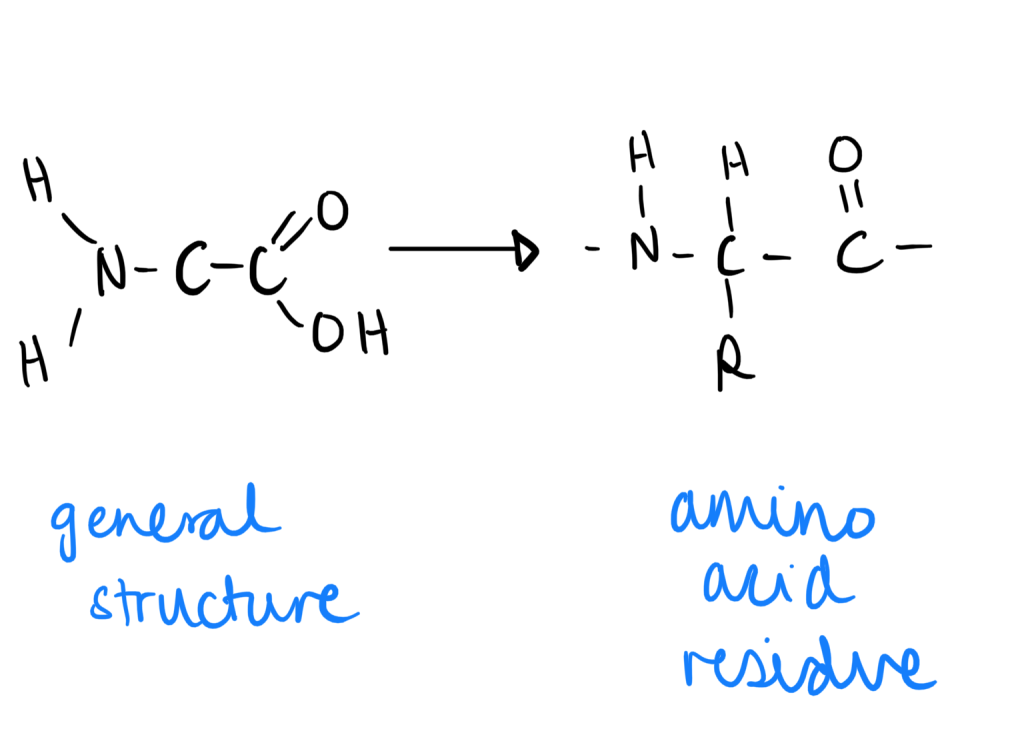
Amino Acid Residue
- When an amino acid is in a peptide bond (through condensation), the remaining portion is called the amino acid residue.
Classification of Amino Acids
- Amino acids are classified through their R groups. Each amino acid has a very specific R group.
- Chemical properties include being acidic, basic, neutral, polar (hydrophillic and forms H bonds) or non-polar (hydrophobic) .
- Physical properties of amino acids include aliphatic or aromatic. An aromatic molecule has a cyclic, planar structure and is highly stable. An aliphatic molecule contains just hydrogen and carbon.
pKa and R Groups of Amino Acids
- The pKa is the pH at which there is no overall charge . It is a measure used to indicate the strength of an acid. The charge of an amino acid (positive/ negative) depends on the pK of the side group.
| Positively Charged amino acids | Negatively Charged amino acids |
| Lysine | Glutamate |
| Histidine | Asparate |
| Arginine |
- If the pH value of the solution < pKR value then the side group will be protonated (addition of hydrogen ions).
- If the pH value of the solution > pKR value then the side group will be de-protonated (loss of hydrogen ions).
- The whole aim of the amino acid is to get the pH close to the pKa. If there is a low pH, there are free hydrogen ions. The amino acid will accept them and become protonated to try and raise the pH closer to the pKa. If the pH is high, there is a low concentration of free H+ ions, meaning the amino acid will donate H+ and deprotonate to get the pH closer to the pKa.
Structure of Proteins
- Primary Structure is the linear amino acid sequence which makes the polypeptide chain.
- Secondary Structure is the folding of the polypeptide backbone due to interactions without R groups. These shapes can be alpha helixes or beta pleated sheets depending on the formation of hydrogen bonds formed within structures.
- Tertiary structure is the 3D configuration. This is based on the bonding within R groups of amino acids.
- Quaternary Structure is the interaction between multiple polypeptide chains and prosthetic groups. These forms protein subunits. For example, haemoglobin is made up of multiple polypeptide chains and a prosthetic group containing Fe2+.
Peptide Bond formation
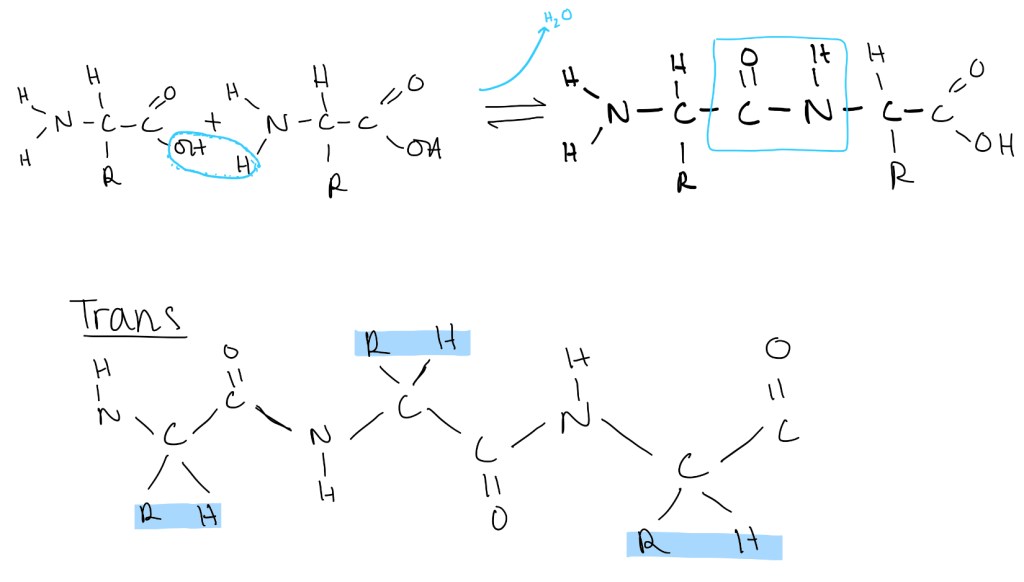
- A link forms between 2 amino acids through a condensation reaction as a peptide bond is formed and a molecule of water is released.
- The peptide bond is planar. The carbon, hydrogen, nitrogen and oxygen atoms that make the general structure are all on the same plane.
- The bond between the carbon and the nitrogen has a partial double bond causing limited rotation. This contributes to the atoms being on the same plane.
- Trans formation is favoured by amino acids. This is where the R groups are on opposite planes from one another. The cis version causes a lot of steric hindrance. This is not favourable for binding and protein folding.
- The bonds on either side of a peptide bond are allowed to rotate. This allows flexibility of the protein and allows the protein to fold into a specific shape according to the amino acid sequence.
Isoelectric Points of Proteins
- This is the pH at which there is no overall net charge amongst the structure of the protein.
- Basic proteins: pI>7 and is positively charged based on the R groups.
- Acidic Proteins: pI<7 and negatively charged based on the R groups.
Conjugated Proteins
- Can be covalently bonded to chemical components alongside amino acids
- Lipoproteins: contain a lipid group which is linked with the protein
- Glycoproteins: contain a carbohydrate covalently attached to the protein
- Phosphoproteins: contain a phosphate group which is covalently linked to the protein
Secondary Structure of a Protein
Alpha Helix
- Polypeptide chain folded in a right handed spiral
- Hydrogen bonds form between the N-H of one amino acid residue and the C=O of another amino acid residue roughly 4 amino acids apart
- Small hydrophobic residues are strong helix formers (e.g. alanine and leucine)
- Ferritin (iron storage protein) – majority of the protein is alpha helix structure.
Beta Pleated Sheet
- The polypeptide chains are very stretched out
- R groups alternate to opposite sides of chains
- Anti-parallel structure is more favourable due to optimal Hydrogen bonding
- Parallel sheets run in the same direction but are less stable
- Mixed patterns are also possible where a sheet can be both parallel and anti-parallel
Tertiary Structure
- The formation of bonds between R groups of amino acids allowing maximum stability to be reached
- Types of forces include hydrogen bonding, ionic bonding, hydrophobic bonding and disulphide bridges
- Ionic bonding forms between negative charges on R groups
- Hydrophobic bonding is formed when non-polar R groups cluster together within the inside of the protein molecule.
- Disulphide bridges form between cysteine and is a covalent bond between sulfur atoms. These are the strongest type and the most stable type that contribute to the tertiary structure of a protein.
Fibrous Proteins
- Have a role of support, protection and shape
- Only has 1 type of secondary structure and usually long strands or sheets
- Example includes collagen
Globular Proteins
- Role of regulation and catalysis
- Contains many different types of secondary structures
- Compact shapes can be formed – hence good for working with a small amount of space
- Examples: Enzymes
Water soluble proteins
- When a protein is folded, the small hydrophobic side chains (non-polar) of the molecule are buried deep within the protein in the middle.
- The hydrophilic chains are towards the outside of the protein- they stick out. These chains tend to be polar. They are on the surface so they can interact directly with water molecules.
Quaternary Structure
- Some proteins are made up of two or more polypeptide chains with additional prosthetic groups. The way in which they interact is called the quaternary structure of the protein.
- Haemoglobin has 4 polypeptide chains or 4 subunits. There are 2 alpha chains and 2 beta chains. There is also an inorganic prosthetic group which is the haem group and contains Fe (II) ions. This Fe binds to the Oxygen and allows the haemoglobin to function.
- Collagen is made up of 3 polypeptide chains with hydrogen bonds that help the chains to wind around each other and form an alpha helix. The hydrogen bonds give strength to the collagen molecule. Collagen molecules form cross links with each other and this allows further strength of the protein. The cross links are also staggered, which provides additional stability to the protein.
Protein Denaturation
- Any disruption to the bonding or forces that hold the protein together causes the protein to denature.
- Heat causes denaturation by increasing the kinetic energy within the molecule. This causes the enzyme molecules to vibrate, hence causing disruption of the peptide bonds.
- Extreme heat may be used as a sterilisation technique to denature proteins in microorganisms, leading to their death.
- Changes in pH causes changes in the ionisation states of the amino acids. This can alter hydrogen bonds (secondary structure) or the ionic bonds (in the tertiary structure) of the proteins.
- Organic solvents have hydrophillic and hydrophobic portions which are attracted to hydrophillic and hydrophobic amino acids present in a polypeptide structure. This pulls the amino acids hence pulling the whole protein structure apart, denaturing the shape.
Misfolding of Proteins
- Misfolding of proteins can occur due to a number of reasons including genetic defects and environmental factors. This can then lead to disease
- In Alzheimer’s Disease, there is an abnormal misfolding of the beta-amyloid protein. This is the broken down product of the amyloid precursor. It’s thought to be toxic in the brain in abnormal levels as the proteins clump together to form neural plaques.
- There is also an abnormal accumulation of tau proteins. This is involved in stabilising the cytoskeleton in neurone cells. The abnormal accumulation prevents the transport system of neurones by preventing the transport of chemicals within neurones and the synaptic communication between neurones. This may affect neurone functionality and lead to neural cell death.
Enzyme Regulation
Regulation of enzymatic activity:
- Short term Regulation
- There are 3 ways of changing enzyme conformation: allosteric regulation, proteolytic cleavage and covalent bonding. These are examples of short term regulation of enzymes.
- Allosteric regulation is the binding of a substance away from the active site of the enzyme causing the tertiary structure of the enzyme to change. This prevents the binding of the substrate to the active site.
- Covalent bonding will change the bonding in the enzyme, altering the shape of the active site
- Proteolytic Cleavage is the removal of an enzyme subunit which causes a change in shape, allowing enzyme to function.
- Long term regulation of enzymes include the regulation of protein synthesis (this is for example post-translational modifications, and this affects the number of enzymes present) and protein degradation (changes rate at which protein/enzyme is broken down).
Substrate and Product Concentration
- Isoenzymes are different forms of the same enzyme. These enzymes catalyse the same chemical reaction. However, there binding sites and binding affinities differ, alongside kinetic properties, amino acid sequence and composition, and electrophoretic properties (this causes difference when enzymes are separated via gel electrophoresis) .
- A coenzyme is an organic non-protein molecule that helps an enzyme’s catalytic activity. These are not enzymes themselves. They can’t function alone. A way to control the activity of an enzyme is by limiting coenzyme availability.
- Enzyme signalling pathways: sometimes a process called product inhibition can inhibit an enzyme pathway. As product amount grows the forward reaction is inhibited. When there is enough product produced a signal is sent which shuts down enzyme activity. Glucose-6-phosphate inhibits hexokinase activity through competitive binding (blocking through binding on the active site) or allosteric regulation.
Allosteric Regulation
- This shows a sigmoidal relationship between rate and substrate concentration.
- The Vmax (maximum rate) remains the same as simple enzymes but the Km (concentration at which 50% of Vmax is reached) differs.
- The enzyme can be in the Tense T state – low affinity, or the Relaxed R state- high affinity.
- When allosterically bound the enzyme remains within the T state making substrate binding and hence protein function difficult.
- Binding of a substrate to one subunit makes complex formation in other subunits a lot easier.
- Allosteric activators increase the proportion of enzymes in the R state, whereas allosteric inhibitors increase the proportion of enzymes in the T state.
- Phosphofructokinase is an allosterically regulated enzyme that paces glycolysis.
- When there is a high amount of ATP present, constant glycolysis will lead to excess ATP and hence needs to be slowed down
- ATP, H+ and citrate ions will act as allosteric inhibitors, which will bind to PFK-1 and limit its activity. A sigmoidal graph is seen.
- When ATP concentration is low, the PFK-1 is not inhibited and a normal enzyme graph can be presented. AMP and fructose-2,6-biphosphate are allosteric activators.
Buffers and Enzymatic Activity
The Henderson-Hassel Balch Equation:
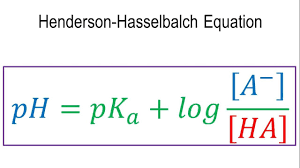
- pKa is known as the acid dissociation constant.
- When acids or bases are added to pure water, H3O+ ions or OH- ions are produced. These in turn increase/ decrease pH.
- Buffer systems are made to reduce large changes in pH. This is done by an already existing A-/HA system. This allows the maintenance of equilibrium
- Added H+ is neutralised by A-, which is converted to HA (acid)
- Added base (OH-) is neutralised by the H+, converted into A-
- The changes in HA/A- ration changes the pH . pH change is related to the ratio of HA/A-, so the change is very small.
Phosphorylation of Proteins
- Adding 2 negative charges on the protein changes the overall charge of the protein.
- Phosphoryl groups can make H bonds. This causes conformational changes and adds stability.
- Rate of phosphorylation/ dephosphorylation can be adjusted, which helps tur off/on a protein.
- Depending on what protein is altered, it can activate or deactivate the protein.
- Energy is required for the phosphorylation as the phosphate group comes when the ATP breaks down.
- Amplification effecs occur when a small activation pathway results in direct activation of other enzymes, introducing an activation cascade.
- Protein kinases transfer terminal phosphate from ATP to the -OH group of Ser, Tyr, and Thr
- Protein phosphatases – reverse effects of kinases by catalysing the hydrolytic removal of phosphoric groups from proteins. This releases the phosphate from the protein , changing conformation of the active site from the protein.
Glycogen Breakdown and Synthesis
- Regulation of glycogen breakdown and synthesis is reciprocal
- Protein kinase A phosphorylates and activates the phosphorylase kinase, which activates phosphorylase A leading to the breakdown of glycogen
- Phosphorylase kinase A also phosphorylates glycogen synthase A leading to the inactivation of glycogen production. This means synthesis can’t occur at the same time breakdown is occurring. It also ensures there isn’t excessive activation of glycogen breakdown.
Proteolytic Cleavage:
- Digestive enzymes are synthesised as zymogens. These are inactive precursors that are proteins. Zymogens have a subunit that will inhibit active site or change conformation in a way that it is an intrinsic allosteric regulator, in the stomach and the pancreas.
| Site of Synthesis | Zymogen | Active Enzyme |
| Stomach | Pepsinogen | Pepsin |
| Pancreas | Chymotrypsinogen | Chymotrypsin |
| Pancreas | Trypsinogen | Trypsin |
| Pancreas | Procarboxypeptidase | Carboxypeptidase |
| Pancreas | Proelastase | Elastase |
- Protein hormones are also examples of inactive precursors as they are initially synthesised
- Blood clotting is stimulated by a series of proteolytic activations to ensure a rapid and amplified response.
- Developmental processes are caused by a series of zymogens specifically for tissue remodelling. The time at which these are activated are specific, however, to avoid inhibition of other enzymes.
- Programmed cell death (apoptosis) is mediated by proteolytic enzymes, caspases, which are synthesised in the inactive procaspase form.
Proteolytic Activation of Chymotrypsinogen
- A digestive enzyme that hydrolyses proteins in the small intestine.
- The precursor is chymotrypsinogen which is synthesised in the pancreas within acting cells. The precursor is stored in membrane bound vesicles i.e. zymogen granules
- Upon stimulation chymotrypsinogen is released into the lumen of the small intestine and activated by another protease trypsin.
- Cleavage (cutting) occurring at different parts of the amino acid sequence of chymotrypsinogen induces changes in proteins, leading to greater breakdown, hence making the protein more relaxed.
- Cleavage occurs at Arg15 and Ile16 generating pi-chymotrypsin (2 amio acid chains)
- Further modification occurs causing pi-chymotrypsin to turn into alpha-chymotrypsin (3 amino acids)
- Cleavage and modification of the protein causes protin to go from the T (tense) state into the R (relaxed) state.
Zymogen Activation and Deactivation
- Activation of pancreatic zymogens is actually regulated by the protein trypsin.
- The trypsin zymogen is trypsinogen
- Activated by the enteropeptidase which is a membrane bound protease in the lining of the duodenum
- Active trypsin can autocatalytically activate trypsinogen. This causes amplification through a cascade, meaning few active trypsin molecules can make a large effect.
- Forms a positive loop where more trypsin presence causes more proenteropeptidase to activate enteropeptidase.
- Trypsin is inhibited by the pancreatic trypsin inhibitor stopping the activity if the enzyme
- Alpha1-antitrypsin is a plasma protein that can inhibit many proteases. When the inhibitor is bound to the enzymes it can inhibit chymotrypsin, elastase, kallikrein, phospholipase A
- The disease emphysema is caused by a lack of alpha1-antitrypsin, resulting in the destruction and disfunction of alveolar walls in the lungs.
Blood Clotting Cascades
- There are two blood clotting pathways: intrinsic and extrinsic
- Intrinsic pathway: caused by the damage of the endothelial lining of blood cells which promotes binding of factor 12
- Extrinsic pathway: traumatic release of tissue factor, factor 3. This comes from tissue surrounding the capillary rather than the capillary itself.
- Both factors lead to the activation of factor X, causing thrombin activation, allowing the clot to form. Very small amounts of factor 3 and 12 also cause blood clotting
- 12 different types of blood clotting factors
- Activation occurs through phosphorylation and proteolytic cleavage of several different factors
- This allows fibrin production, which caused blood clots to become hard.
Prothrombin Structure and Function
- Protease function (thrombin) is within the C terminal domain
- The ‘kringle’ domains allow the thrombin to remain within its inactive form.
- Gla domain target it to certain sites of activation. Gla residues are formed through the addition of carboxyl group to glutamate residues in order to form carboxyglutamate
- Prothrombin binds Ca2+ ions to activate, if there is no carboxylation of glutamate then you dont get activation of thrombin, no activation of fibrin activation of thrombin, no fibrin, no clot
- The kringle and Gla domains need to removed in order to allow the activation to occur
- Prothrombin activation is specific. Only the prothrombin that is in specific to the place that requires a blood clot (sites of damage) is activated.
Fibrinogen
- The structure has 2 sets of tripeptides: alpha, beta and gamma. These are all joint at the N-termini.
- Thrombin cleaves fibrinopeptides A and B from the central globular domain of fibrinogen
- Globular domains at the C ends of the beta and gamma chains interact with the exposed N-terminus of the cleaved fibrinogen A and B chains. This allows the mesh clot to form.
- Factor 12 allows the cross-linkage to form between the chains of the clot. This is done by transglutaminase and protransglutaminase.
- These enzymes rely on glutamine. The glutamine is modified by the above enzymes that allow the glutamine to bond with lysine in the fibrin chain to form the mesh network.
Haemophilia
- Haemophillia A is cuased by a defect in factor VIII causing less thrombin activation.
- Haemophillia B is caused by a deficiency in factor IX, leading to effects in the intrinsic pathway. These are less severe than the consequences of Haemophillia A
- Recombinant factor VIII is used treat as it counteracts the defected factor VIII allowing the clotting cascade to continue.
- Factor VIII leads to the activation of Factor IX. Hence it is a coenzyme, not a protease.
Positive Feedback with Fibrin
- Constant activation of the clotting pathway is needed for the blood clot to form
- Positive feedback loop allows Factor XI, V and VIII and thrombin to activate. This causes greater fibrin production.
- Without the positive feedback loop, the blood clot would be weaker, hence not sufficient enough for its function.
Inhibition of the blood clotting cascade
- Blood factors dissolve into the blood flow and are processed and removed by the liver
- Digestion occurs through proteases:
- Factor Va and VIIIa are degraded by protein C
- Protein C is activated by thrombin binding to an endothelial receptor
- Defects cause thrombotic diseases
- Protein C knocks factors and allows them to degrade
- Specific inhibitors include Warafin and antithrombin III (AT3) which binds to thrombin and prevents thrombin from being produced. This is enhanced by heparin binding.
Control Points in Blood Clotting
- Inactive zymogens are present at low concentrations
- These zymogens become cleaved, through proteolytic activation
- Amplification of initial signal by a cascade mechanism
- Clustering of clotting factors at the site of damage
- Feedback activation of thrombin and fibrin allows continuous clotting to occur, strengthening the clot.
- Termination of clotting occurs through multiple mechanisms
- Clot breaks down by proteolytic activation.
Catabolism
Overview
There are 4 main stages of catabolism:
| Stage | Events |
| Stage 1 | Breakdown to Building Block molecules |
| Stage 2 | Breakdown to metabolic intermediates, releases reducing power and energy (only a small amount) |
| Stage 3: Kreb’s Cycle | Inputs: 2 Acetyl CoA, 6 NAD+, 2 FAD, 2 ADP + 2P Outputs: 4 CO2, 6 NADH, 2 FADH2, 2 ATP Provides reducing power and energy for oxidative phosphorylation (NADH, FADH2 and ATP) |
| Stage 4: Oxidative Phosphorylation | Inputs: 10 NADH, 2 FADH2, 6 O2 Outputs: ATP (26-28 per glucose), 6 H2O, NAD+, FAD Completes the conversion of reducing power (FADH2 and NADH) into energy currency in the form of ATP |
Stage 1 Catabolism:
- Converts nutrients to a form that can be taken up by cells.
- Proteins get converted to amino acids
- Carbohydrates get converted into monosaccharides
- Lipids get converted into glycerol and fatty acids
- Alcohol is already in a form that can easily be taken up by cells
- Occurs in the GI tract, extracellular
- No energy is produced in this stage. It involves the breakage of C-O bonds and C-N bonds, but only phosphate bond breakage produces a high amount of energy.
Stage 2:
- Breakdown of small molecules into even smaller molecules
- Amino Acids = NH4+, further breaking down into urea, alpha-keto acids, pyruvate, acetyl coA
- Monosaccharides breakdown into pyruvate
- Glycerol breaks down into pyruvate
- Fatty Acids breakdown into Acetyl CoA
- Alcohol also breaks into Acetyl CoA
- Occurs intracellular in the cytosol and the mitochondria
- Doesn’t occur in all cells. RBC don’t have any mitochondria hence only rely on glycolysis
- Stage is oxidative – they require co-factors that get reduced
- Some energy is produced in the form of ATP – 2 molecules of ATP per glucose
- Humans cant survive off glycolysis alone, need oxygen for ETC and oxidative phosphorylation to produce sufficient ATP
Stage 3:
- Involves The Citric Acid Cycle, Kreb’s cycle
- Inputs: 2 Acetyl CoA, 6 NAD+, 2 FAD, 2 ADP + 2P
- Outputs: 4 CO2, 6 NADH, 2 FADH2, 2 ATP
- Occurs in the mitochondria
- Oxidative, requires NAD+, FAD
- For every molecule of glucose, 32 molecules of ATP is
- produced via Kreb’s cycle
- Carbon ends up as carbon dioxide, acetyl Ch3CO is converted to 2CO2
- Useful for producing precursors for anabolism due to various different inputs, not just acetyl CoA
Stage 4:
- Involves the electron transport chain and ATP synthesis via oxidative phosphorylation
- Electron Transport chain + Oxidative Phosphorylation :
- Inputs: NADH, FADH2, O2
- Outputs: NAD+, FAD, ATP, H2O – NADH and FADH2 are re-oxidised
- Oxygen is reduced to H2O
- Oxygen is essential for ETC, if it is not present, the chain does not work, no oxidative phosphorylation and no ATP produced
Carbohydrates
- General Formula of (CH2O)n where n is the length of the chain
- Only consists of carbon and oxygen groups
- Contains aldehyde (CHO) and ketone groups (C=O)
- Contains multiple OH groups
- Monosacchardies = simple single sugar units (3-9 C atoms)
- Further categorized as aldose or ketone groups:
- Aldose: aldehyde-containing sugar – Ketose: keto-containing sugar
Polymers of monosaccharides:
- Disaccharides: 2 units (lactose, sucrose, maltose)
- Oligosaccharides: 3-12 units (dextrins)
- Polysaccharides: 10 – 1000 units (glycogen, starch cellulose, all polymers of glucose)
- Varies according to the length of the carbon chain within them
More about monosaccharides:
- Triose: 3 carbons
- Pentose: 5 carbons
- Hexose: 6 carbons
- Monosaccharides with 5 or more carbon atoms have ring structures. They can be alpha or beta rings
- Alpha rings have the OH group on the first carbon position on the opposite side of CH2OH group
- Beta rings have the OH group on the same side as the as the CH2OH group
- Rings are polar and not fat soluble and are very hydrophilic. For this reason they need transporters to travel across the membrane. They are also partially oxidised so do not need further oxidation.
Disaccharides:
- Formed by the condensation of 2 monosaccharides
- Form oxygen gylcosidic bond
- Water is eliminated in bonding
Glycogen:
- Used as a storage molecule. Stored within the liver and the skeletal muscle
- Condensed into and stored as granules
- Can be stored as a large chain or a branching structure, leading to a branched molecule
- Branching means there are many attack locations for enzymes to breakdown glycogen, and to free up glucose molecules very quickly
- Branching also prevents the binding of water molecules so we don’t need to store too much water.
Starch
- Another polysaccharide found in plants
- Enzyme amylase breaks down starch into glucose and maltose
- Digestion of starch usually occurs in the GI tract
Cellulose
- Also another polysaccharide found in plants
- Can’t be digested by humans – lacks the GI enzymes for digestion of cellulose
- Good for dietary fibre
- Adds to peristalsis and provides roughage (fibre that aids in travel of food through the gut)
Metabolism of Carbohydrates
- Breakdown of carbohydrates involves the breakage of glyosidic bonds to monosaccharide units glucose, galactose and fructose mainly
- Saliva breaks down starch in the mouth using amylase and glycogen using the enzyme dextrin
- Pancreas breaks down carbohydrates using amylase to form monosaccharides
- Small intestine also contains enzymes for the breakdown of carbohydrates: lactase for lactose, sucrose for sucrose, pancreatic amylase for alpha 1-4 bonds, isomaltase for alpha 1-6 bonds
Absorption
- Monosaccharides are actively transported into the intestinal epithelial cells and then via blood supply to target tissue where they enter cells.
- Enters the bloodstream then goes to the liver via the hepatic portal duct. The liver then breaks down the simple smaller molecules
- First pass effect: whatever is broken down, digested and absorbed goes to the liver. This is important as this is the way that drugs work: they get broken down and sent to the liver.
- These nutrients are then taken up by GLUT1 and GLUT2 transport proteins into cells via facilitated diffusion.
- Proteins can be regulated by hormones. This can be such as insulin where GLUT4 is stimulated to take up glucose from the blood to reduce blood sugar levels.
Pharmacodynamics
- Pharmacodynamics is all about the mechanisms and effects of medications on the body.
- Drugs always bind to receptors. Receptors are specialised forms of proteins that can be intracellular or extracellular and bind to a molecule also known as a ligand. This forms a complex.
- A ligand can be a neurotransmitter, hormone, or even a drug.
- The receptor-ligand complex gives rise to a signalling cascade of intracellular molecules. These are known as secondary messengers. Once this signalling cascade is activated it causes a change in function.
- Intracellular receptors can be in the nucleus or the cytoplasm. These bind to hydrophobic ligands, such as steroids, which can easily diffuse through the phospholipid bilayer. Once the complex has formed, they bind to specific parts of DNA which can activate or inhibit certain genes.
- Extracellular receptors are on the cell surface membrane. They tend to bind to hydrophilic molecules. There are 3 main types of extracellular receptors.
- First type of an extracellular receptor is the ligand-gated ion channels. Normally, these ion channels tend to be closed.
- Once a specific ligand binds and forms a complex, the ion channels will open and selectively allow certain molecules to pass through into the cytosol. Examples include passive diffusion of Na+, Ca2+, K+ and Cl-, all entering in order to activate a signalling cascade within the cell.
- The second type of extracellular receptor is the G-Protein Coupled Receptors. This is a seven transmembrane long protein, with an N-Terminus that sticks out extracellular and the C-Terminus which sticks inwards, intracellular.
- The N-terminus will bind to the ligand and form a complex. The C-terminus then attaches to G-proteins (guanine nucleotide binding proteins). These contain an alpha, beta and gamma subunit. Usually the alpha subunit binds to a GDP molecule. In this state the G-protein is inactive. Once a complex has formed with a ligand, the G-protein experiences a change in shape which causes the exchange of GDP with GTP. This detaches the alpha subunit, which transports to activate the signalling pathway. There are 3 types of G-Proteins (Gq, Gi and Gs). Each type has its own alpha subunit hence they cause different changes in function.
- Gq tends to activate phospholipase C. This cleaves a type of phosphate. Gs and Gi stimulate or inhibit adenylyl cyclase. This takes ATP and transforms it to cAMP. These then activate different sets of enzymes and molecular pathways.
- Another type of extracellular receptor is the enzyme coupled receptor. This binds to a ligand and forms a complex on the extracellular end. On the intracellular end there is an an enzymatic domain that is usually a protein kinase.
- This enzyme tends to phosphorylate other molecules. As the complex forms, a conformation change occurs to the enzymatic domain. This provides a high affinity for secondary messenger molecules and proteins to bind to the enzymatic domain. These get phosphorylated by tyrosine kinase and activate a signalling cascade.
- Each medication is specific to a type of receptor molecule. This is like the lock ad key model. This is a mechanism that can be used to explain drug specificity. However, drugs can never be fully specific to only a certain type of receptor- they can also bind to other receptors which are very similar. This is why drugs have side effects.
Key Pharmacodynamic Terms
- Affinity is how strongly a drug can bind to a receptor. This depends on the chemical bonds between the drug and its receptor.
- Potency is described as the amount of medication needed to produce an effect on an individual. (A higher potency means less of it is needed to cause an effect)
- Affinity and potency are in a positively linked relationship – the higher the affinity, the higher the potency, hence less of that drug is needed to allow treatment of a patient.
- Intrinsic activity is the ability of a drug to activate a receptor. This is of course also essential for the drug to cause an effect, as that is how it will alter body function.
- The intensity of response towards a medication depends on the intrinsic activity and the number of receptors that are bound to the drug.
- The efficacy of a drug is the maximal effect that the drug can produce as part of its effectiveness.
Pharmacodynamics Graphs
- A graph can be produced of dosage/ concentration (x axis) against response (y axis).
- This produces a sigmoidal curve, like an S shape. At the beginning the curve is quite flat. This is because not that many receptors bind, since thee aren’t that many ligand drugs. As the graph goes through a steep increase, this is because more drug molecules are present to bind to receptors and form a complex. At the end as the graph levels off again, this is because all receptors are occupied and adding more drug won’t make a difference.
- This is known as Emax – the maximum binding capacity of the drug receptors. This produces the maximal response in turn.
- EC50 is the described as 50% of the maximum effect that is produced. This can be used to describe potency. If EC50 is lower, less of the drug is needed to produce that response, hence it has a higher potency.
- EC50 and potency have a negatively linked relationship – the lower the EC50 value, the higher the potency. This is why the more left the value of EC50 is on a dose-response curve, the higher the potency.
Membrane Transport and Ca2+ Regulation
- Cellular Membranes are semi-permeable: they allow some substances to pass through but not others.
- The cell membrane is composed of a phospholipid bilayer and membrane proteins. The permeability of a molecule therefore depends on its ability to pass through the phospholipid bilayer or the membrane proteins.
- The phospholipid bilayer allow both small, non-polar molecules to pass through as well as hydrophobic molecules to pass through as well. Examples include O2, CO2, water, urea and glycerol.
- In order to allow larger and charged particles to pass through the membrane, this is dependent on the expression of protein channels and transporters. This depends on the type of cell.
- Neurone cells, for instance, have a higher permeability to Na+ and K+ ions in comparison to normal cells due to the generation of an action potential.
Passive Transport
- Passive Transport is via diffusion.
- This occurs down an electrical or chemical gradient without requiring an input of energy.
- Diffusion through the bilayer itself is called simple diffusion, but diffusion occurring through membrane and transport proteins is called facilitated diffusion.
- Facilitated diffusion is a saturable process (reaches a maximum rate) due to the use of channel and membrane proteins, as these can eventually become limited in number.
- Some channels are only open when certain substances are present. These channels are called gated channels. These include ligand gated channels, voltage gated channels and mechanically gated channels.
- Ligand gated channels are channels that are only active when binding to a ligand (a chemical signalling molecule)
- Voltage gated channels are channels that open in response to changes in membrane potential (e.g. depolarisation)
- Mechanical receptors are active when there is a stretch sensation in the cell membrane. This is for example mechanoreceptors in the skin.
Electrochemical Gradients
- Whether transport is passive or active depends on the electrochemical gradient. If there is a gradient then the particles will diffuse passively however if there isn’t a gradient the particles will diffuse actively.
- Free energy change can determine whether a substance diffuses passively or actively. If the free energy change is negative (-ΔG), diffusion will occur passively however if free energy change is positive (+ΔG), diffusion will occur actively.
- Formula for uncharged particles: we only consider the concentration gradient.
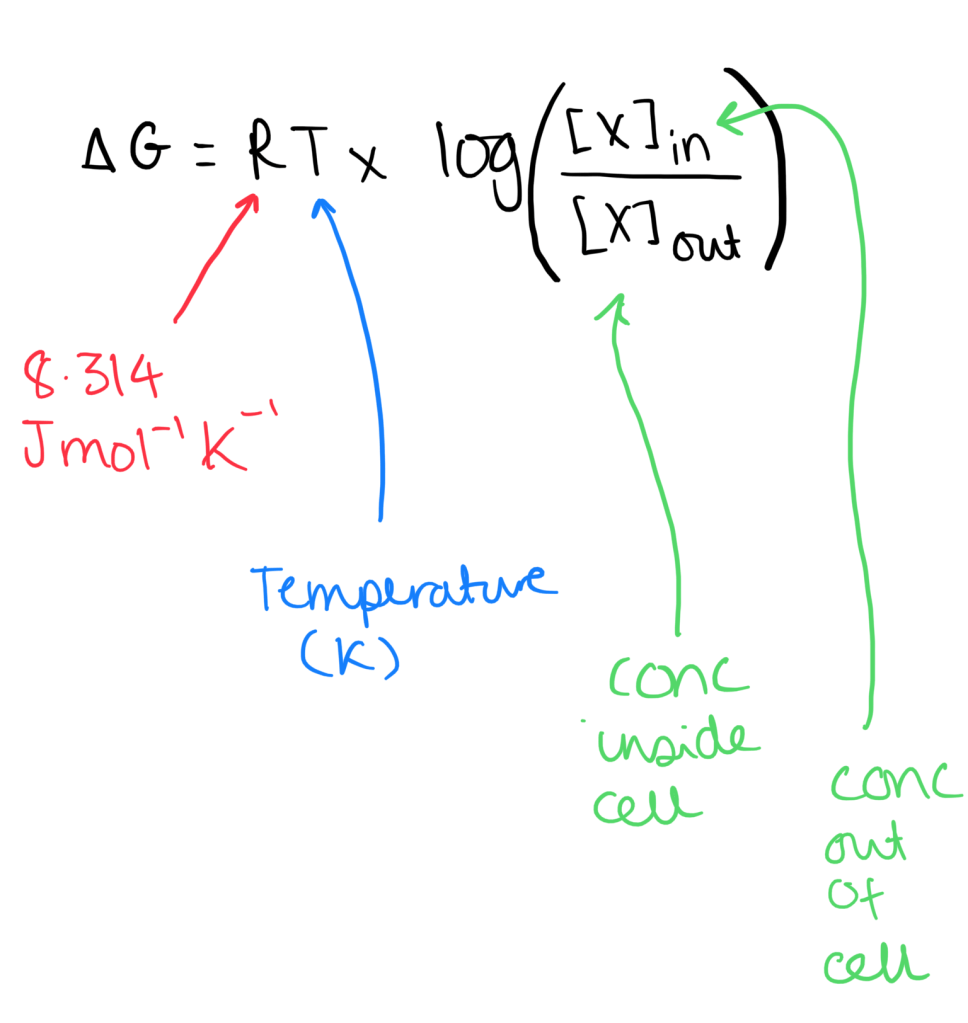
- Formula for ions – since they are charged we have to observe the electrical gradient (this is the membrane potential).
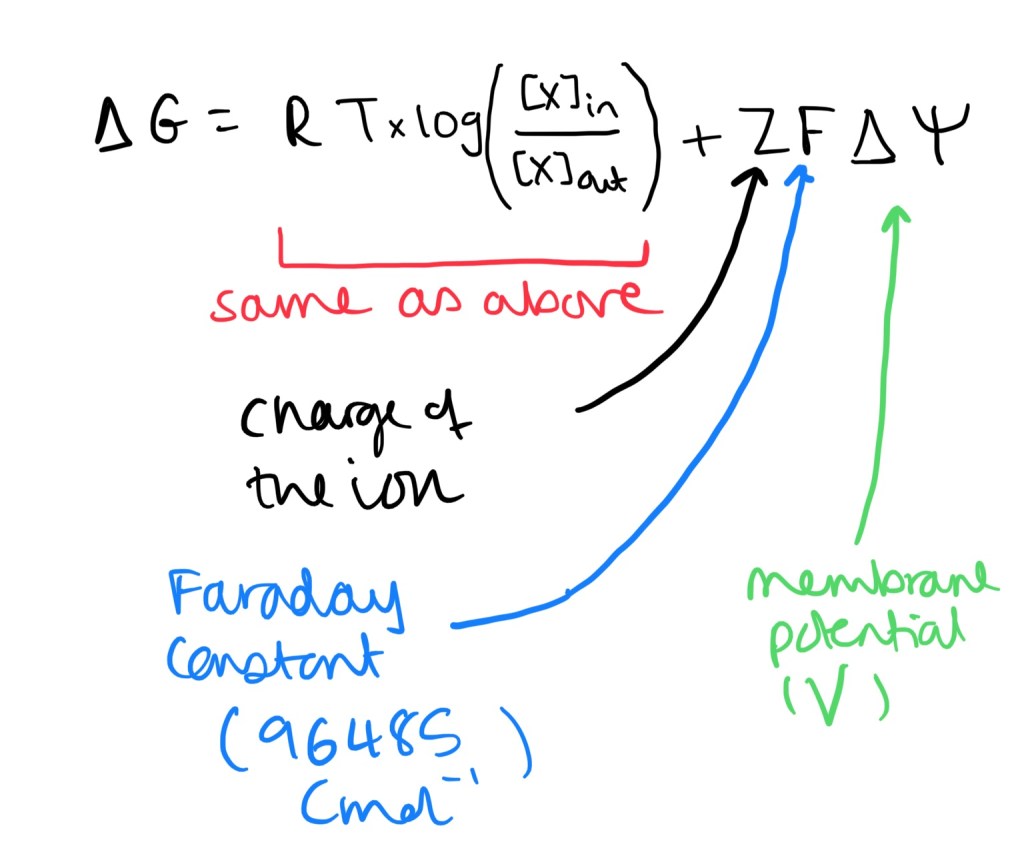
Active Transport
- This is the movement of ions and molecules against an unfavourable electrochemical gradient, hence requiring energy via ATP hydrolysis for the transport to occur.
- Active transport requires a lot of energy. Up to 50% of the cells metabolic rate is derived from energy through active transport.
- Primary active transport is where energy is directly direvied from ATP hydrolysis. This is achieved through pump proteins, or ATPases. This includes the Na+/K+ ATPase, or the plasma membrane Ca2+ ATPase.
- Secondary active transport is also known as co-transport. This uses energy from a pre-existing chemical gradient, such as Na+ from the Na/K ATPase.
- Co-transport: symport: such as the Na+-glucose cotransporter. This is where the ions and molecules move through the co-transporter in the same direction.
- Antiports such as Na+/H+ co transporters are where the molecules move in opposute directions.
Cellular pH and Volume Regulation
- Intracellular pH is managed through the movement of H+ (acid) or HCO3- (base).
- Na+/H+ ATPase. This extrudes H+ ions, increasing the intracellular pH. It uses a pre-existing gradient to move 1 Na+ into the cell and 1 H+ out of the cell (antiport). This is a type of secondary active transporter. (Kidney Tubules)
- The Na+/HCO3- transporters causes an alkali influx. It uses a pre-existing Na+ gradient to move 1 Na+ into the cell and 3 HCO3- into the cell. (Many different types of cells).
- The anion exchanger acidifies the cell. It moves 1 Cl- into the cell and one HCO3- out of the cell. (Found on RBC and Kidney Tubules)
- Cell volume regulation is the movement of osmotically active ions into or out of cells, and this as a result causes the movement of water into and out of the cell. An influx of ions can cause an influx of water, whereas an efflux of ions causes an efflux of water.
Intracellular Calcium Regulation
- Ca2+ ions is involved in a range of intracellular signalling pathways.
- It may help to regulate physiological and pathological processes such as fertilisation, muscle contraction, neurotransmission, metabolic regulation, cell death, learning and memory.
- Extracellular concentration of soluble is much higher. This can be found in blood or interstitial fluid, and half is attached to plasma proteins such as albumin.
- Concentration of Ca2+ intracellular is extremely low as a high concentration may even be toxic to the cell. The endoplasmic reticulum (or the sarcoplasmic reticulum in muscles) is a good storage space for Calcium ions.
- Extracellular concentrations of Ca2+: 1 x 10^-3 M
- Intracellular Concentrations of Ca2+: 1 x 10^-7 M
- ER/SR Concentrations of Ca2+: 1 x 10^-4 M
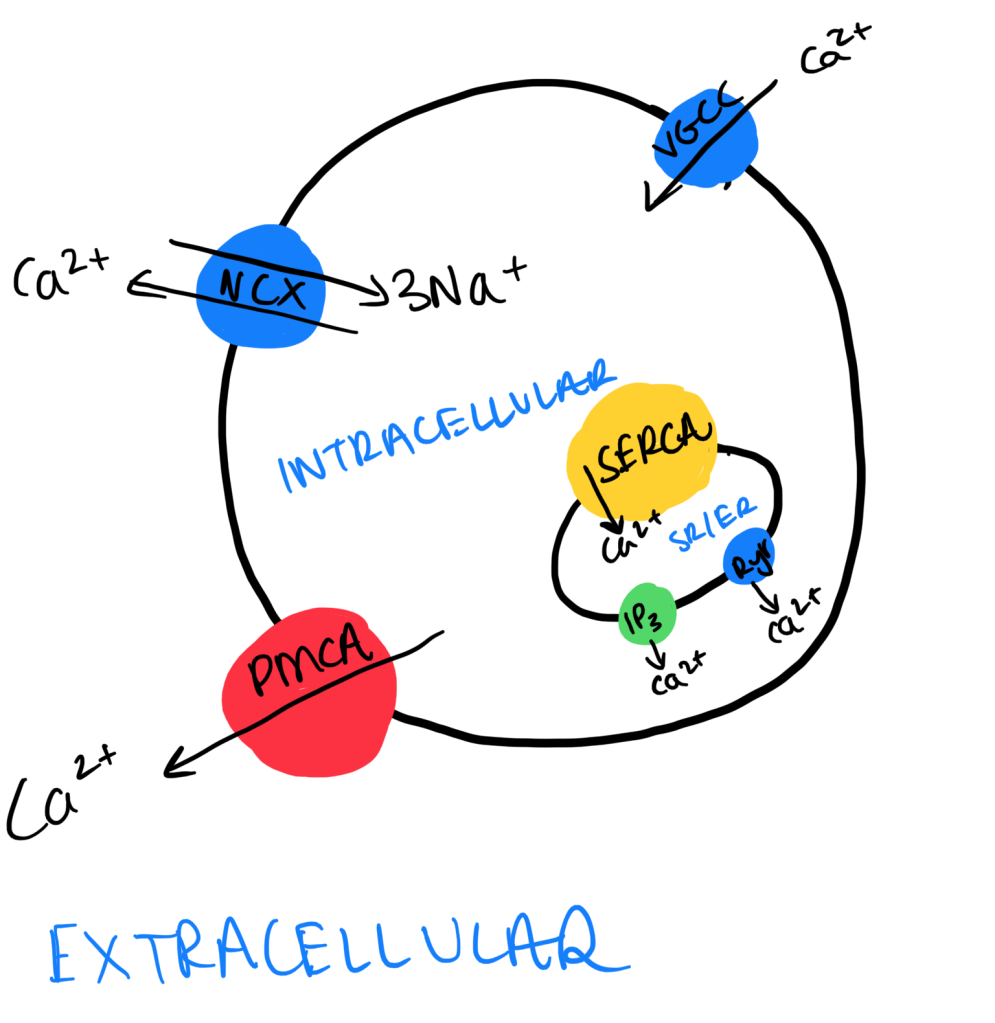
Regulation of Ca2+ through protein transporters:
- Na+/ Ca2+ Exchangers (NCX): Usually found on the plasma membrane. It usually takes 3Na+ into the cell and 1 Ca2+ out of the cell. Low affinity and high capacity means it is the main Ca2+ extruder.
- Plasma Membrane Ca2+ ATPase (PMCA): Moves Ca2+ out of the cell using ATP hydrolysis. This has a high affinity and low capacity, hence does a small proportion of Ca2+ extrusion.
- SR/ER Ca2+ ATPase (SERCA): Takes Ca2+ out of the SR/ER by ATP hydrolysis. Also has a high affinity and a low capacity.
Rise in intracellular Ca2+:
- Voltage-gated Calcium Channel (VGCC): This protein causes a Ca2+ influx in reponse to the depolarisation of the plasma membrane.
- IP3 Receptor (IP3R): A ligand gated ion channels also found on the SR/ER. The ligand is IP3, generated by GPCRs coupled to G-alpha-q, in turn activating phospholipase C. IP3 binding causes opening of Ca2+ channels and results in movement from the SR/ER to the cytoplasm.
- Ryanodine Receptor (RyR): This is another ligand gated channel found on the SR/ER. This is involved in the process of calcium-induced-calcium-release (CICR), where the binding of Ca2+ to Ryr results in the opening of Ryr and an increased efflux of Ca2+ from the SR/ER. This process is used to increase the amplification of Ca2+ secretion – the extra Ca2+ is derived from the CICR.